 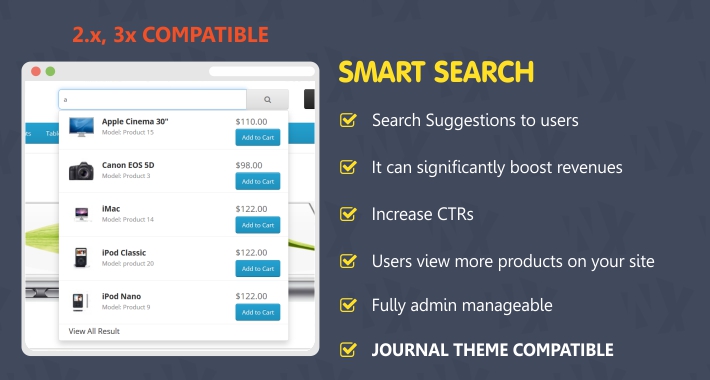
Due to the above point they can be trained from scratch on domain specific texts.They are ‘lightweight’ when compared to transformer models in all areas that matter when creating scalable services (model size, training times, inference speed).However they are still relevant today for the following reasons: Since the introduction of sophisticated transformer models like BERT, word vector models can seem quite old fashioned. Create word vectors build a fastText model Running on a Colab notebook, this can process over 1,800 notices a second. The above code splits our documents into a list of tokens whilst performing some basic cleaning operations to remove punctuation, white space and convert the text to lowercase. Superfast searching of our results using the lightweight and highly efficient Non-Metric Space Library (NMSLIB).We will still be using this algorithm to power our search but we will need apply this to our word vector results. We will train a model on our data set to create vector representations of words (more information on this here). In order to achieve this, we will need to combine a number of techniques: Handle spelling mistakes, typos and previously ‘unseen’ words in an intelligent way. Be orders of magnitude faster than our last implementation, even when searching over large datasets.Be able to scale up to larger datasets (we will be moving to a larger dataset than in our previous example with 212k records but we need to be able to scale to much larger data).Be location aware understand UK postcodes and the geographic relationship of towns and cities in the UK.Return relevant results to a user even if they have not searched for the specific words within these results. 
This post will describe the process to do this and also provide template code to achieve this on any dataset.īut what do we mean by ‘smart’? We are defining this as a search engine which is able to: In this post, we want to go beyond this and create a truly smart search engine. If you for whatever reason don't want/can't use cookies, access SMART through this page.In the first post within this series, we built a search engine in just a few lines of code which was powered by the BM25 algorithm used in many of the largest enterprise search engines today. Information about your selected mode is stored in a browser cookie. Normal modeĬlick on the images above to select your default mode. Different color schemes are used to easily identify the mode you're in. Remember you are exploring a limited set of genomes, though. The numbers in the domain annotation pages will be more accurate, and there will not be many protein fragments corresponding to the same gene in the architecture query results. If you use SMART to explore domain architectures, or want to find exact domain counts in various genomes, consider switching to Genomic mode. The protein database in Normal SMART has significant redundancy, even though identical proteins are removed. 
The complete list of genomes in Genomic SMART is available here. In Genomic SMART, only the proteomes of completely sequenced genomes are used Ensembl for metazoans and Swiss-Prot for the rest. In Normal SMART, the database contains Swiss-Prot, SP-TrEMBL and stable Ensembl proteomes. You can use SMART in two different modes: normal or genomic.The main difference is in the underlying protein database used. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
